The Folding@home team released its software to the public in September 2000. The project aims to simulate protein dynamics, including the process of protein folding and the movements of proteins implicated in a variety of diseases such as Cancer, Alzheimer’s, Parkinson’s, Huntington’s, Influenza, and many others. To carry out these simulations you need enormous computing power, which is extremely expensive. Folding@home brings together citizen scientists who volunteer to run simulations on their personal computers. Therefore, they are looking for volunteers who will use their unused computer power to carry out the simulations; if you volunteer you can help them to speed up the process and achieve results more quickly.
Insights from this data are helping scientists to better understand biology, thus providing new opportunities for developing therapeutics.
How can this project help me?
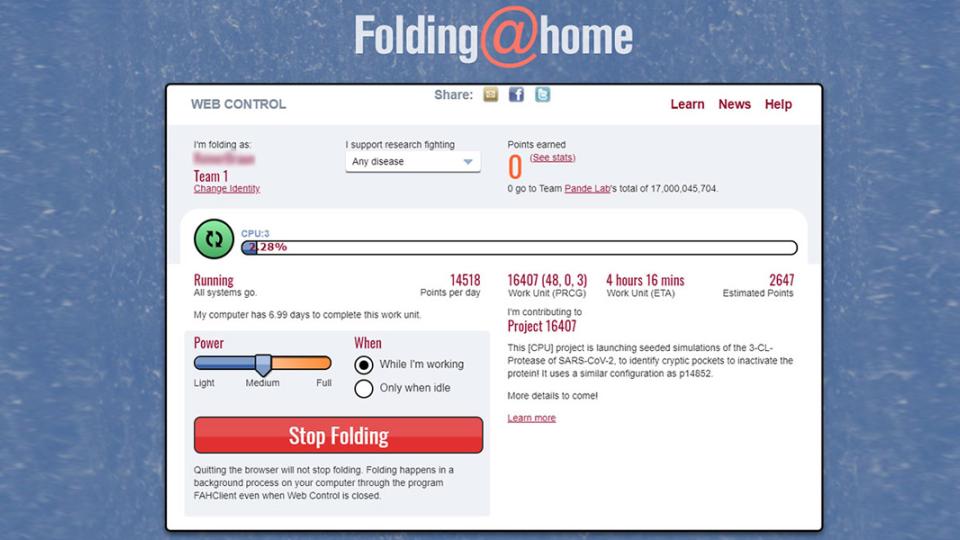
You can learn new things about protein dynamics and how these are related to diseases by becoming part of this citizen science project. You can choose whether to participate alone or in a team, or you can start your own team. The donor and team stats are updated every hour.
How can I use folding@home and what are its benefits?
On their Website, you can download their Folding@home software for free and start helping them to develop/ find cures by simply sharing your computing capacity by running the simulation on your own personal computer. You´ll be rewarded with points depending on the performance of your hardware.